Program nest_abundV1.x
This program computes estimates of parameters described in:
Péron, G., J. Walker, J. Rotella, and J.D. Nichols,
Beyond Survival: Abundance Estimation for Nest Studies
(in prep.)
Abstract
A recurrent issue in studies of bird fecundity is that nests are difficult to find.
Hatching success and fledging success can be inferred from detected nests, but the total number
of offspring per unit of habitat remains unknown. Here we develop a capture-recapture approach
to the issue that deals with imperfect detection probability as well aswith the impact of
environmental and individual covariates on nest fate. We tailor the approach to the estimation of
duck productivity in the Prairie Pothole Region of North and South Dakota. In this study case,
we use the age of the nests at first detection (estimated by the candling method) to make
inference about detection probability. We describe the maximum likelihood estimation of the
total number of nests on the study plots. We find that nesting stage (egg-laying or incubation)
markedly influences both survival and detection probabilities, probably because hens often do
not attend nests when they are not incubating. 6%of the nests were missed by the field crews
during surveys, or failed before the nearest survey, or were initiated after the last survey. This
proportion is expected to be larger in less intense, more typical sampling designs.
nest_abundV1.x was written for a specific dataset, but
may be used for any similar dataset with similar covariates.
Requirements:
- R software (www.r-project.org)
- R package fastGHQuad
- R package mvtnorm
- CSV datafile containing...
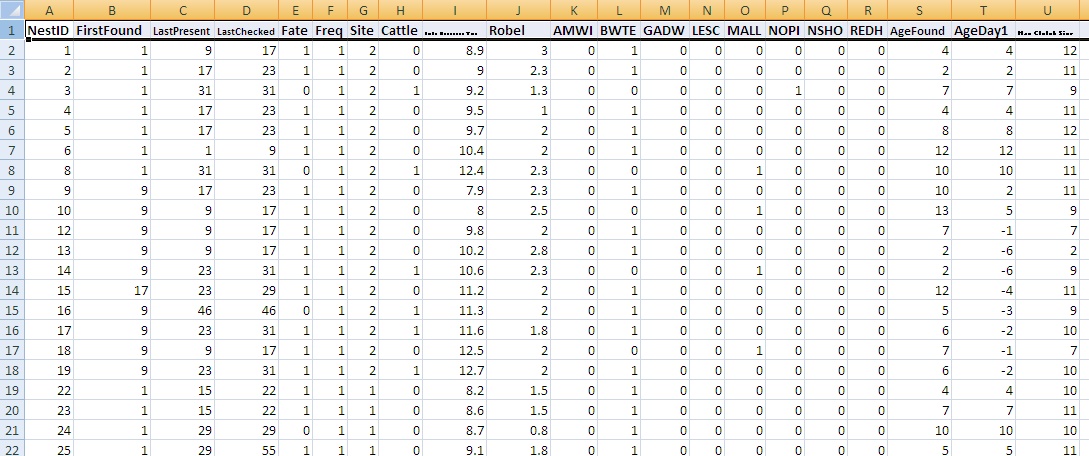
- One row per nest, first line contains field names as listed below:
- The following fields (columns)...
- NestID (numeric or text identifying nest
- FirstFound (day number when nest was first found
- LastPresent (day number when nest was last located
- LastChecked (day number when nest was last checked
- Fate (fate=1 in nest successfully produced young, 0 otherwise
- Site (site group number of nest
- BinCov (Binary covariate for an attribute of the nest.)
- BinCov=1 or 0, depending on the presence or absence of the covariate.
In the sample data, this is 'Cattle', indicating the presence
of cattle near the nest.
- ContCov (Continous covariate for an attribute of the nest.)
- This covariate can be any value between plus or minus
infinity, but we recommend that these be 'normalized' or
'scaled' to prevent optimization problems. In the sample
data, this covariate corresponds to 'Robel'. Since this variable
only ranged from 0.5 to 6.3, they were not normalized or scaled.
- Species1 (species1=1 if nest is occupied by species 1, 0 otherwise)
Species1 corresponds to AMWI in the sample data.
- Species2 (species2=1 if nest is occupied by species 2, 0 otherwise)
Species2 corresponds to BWTE in the sample data.
- Species3 (species3=1 if nest is occupied by species 3, 0 otherwise)
Species3 corresponds to GADW in the sample data.
- Species4 (species4=1 if nest is occupied by species 4, 0 otherwise)
Species4 corresponds to LESC in the sample data.
- ...
- AgeFound (age of nest in days when first found. Age determined
by 'candeling' or other method.)
- AgeDay1 (age of nest on first day of survey)
- Max Clutch Size (maximum size of clutch)
- Note:
- 1st row must contain names specified exactly as listed above
(ie., spelled the same, same upper/lower case).
- Species names mustbe 4 upper case characters.
- Columns D, F, and I in sample data are not needed.
Program operation
- First, create input file similar to the provided sample
dataset, sample_data.csv.
- Download and install R software (available
here)
Note: The R software is available on many web-servers around
the world, so this link will present you with a list of these
servers (or 'mirrors' of the original site). Scroll down the list
to find one close to you, and click it. That site should show
links to download the version of R appropriate to your
operating system.
- Download nest_abundV1.x.zip program (available
here)
- Extract all files from the downloaded zipfile (nest_abundV1.x.zip)
to a folder (eg., My Documents/nest_abund,
Documents/nest_abund, or /home/yourname/nest_abund). Most current
operating systems will allow you to 'unzip' the file simply by
double-clicking it. Older operating systems may require that
you download and install a separate utility program (eg.
7-zip))
to extract the program
files.
- Start R software (Start menu or desktop button)
- Run nest_abundV1.x program (click 'File' menu in R window,
then click 'Source R code' and navigate to the folder where
the nest_abundV1.x was extracted.) Note: The procedure for
starting the program may change when the program is converted
to a 'package'.
- Once nest_abundV1.x starts, it will ask to install
an R package (mvtnorm) if it is not already installed. Click
'yes' to install the package.
- Click the 'Input file' button, then find the input file
you created (or try the provided sample_data.csv file).
- Select the binary covariate (if you have one) by clicking the
BinCov menu and clicking the checkbox next to the covariate name.
In the sample data (sample_data.csv), the binary covariate is 'Cattle'.
- Select the continous covariate (if you have one) by clicking the
ContCov menu and clicking the checkbox next to the covariate name.
In the sample data (sample_data.csv), the continous covariate is 'Robel'.
- The program is configured for a particular study on
Blue-Winged Teal by default. For
other species or study designs, you may need to change
some configuration parameters. This can be done in the
'config' menu. Here are the parameters which can be altered:
- maxbs - number of bootstrap simulations for estimation
of conf. intervals
- maxit - maximum number of iterations for the optimization routine
- GHnodes - number of nodes in Gauss-Hermite quadrature
- LEN3OCC - number of days which include at least 3 occasions
- AvgClutch - Avg. time for clutch completion (=clutch size for
species that lay 1 egg per day)
- Select models you would like to run by clicking the
checkbox next to each model name.
- Click the 'Run' menu, then the 'models' entry.
- Click any model name to view the results
of the associated model (in a Notepad window).
- Quit program using the 'File' menu, 'Exit' selection.
Model names
Model names in this program follow the convention used in other
animal estimation software making it 'easy' to know the
structure of a model from it's name. The models in the program
contain two sets of parameters, one dealing with nest survival
(Phi), and one dealing with detection probability (p). Each
model name was generated by listing each of these two parameters
followed by a list of covariates affecting each one in
parentheses. For example, the model name, Phi(),p(), contains
only two parameters which do not vary according to any covariates.
The model, Phi(site),p(s), produces site-specific estimates
of nest survival (one survival rate for each site), and
stage-specific estimates of detection (stage is abbreviated as
's' in all models, and there are only 2 stages: egg-laying and
incubation). Here is a full list of models and descriptions:
-
Phi()p() - survival and detection are constant across all covariates.
-
Phi()p(s) - survival is constant across all covariates, detection
is potentially different for the two stages.
-
Phi(s)p(s) - survival and detection vary by stage.
-
Phi(site)p(s) - survival varies by site (one estimate per site)
and detection varies by stage.
-
Phi(site+s)p(s) - survival varies by site and stage in 'parallel'
(two estimates per site, egg-laying and incubation, but the
incubation estimates differ from the egg-laying estimates by
a fixed amount (on a transformed, logit scale).
Detection probabilities vary by stage.
-
Phi(site+s+CCOV)p(s) - survival varies by site, stage and the
continous covariate (ccov) in 'parallel'. With a continous
covariate, each nest with a different value for the covariate
can have a different survival rate, but the survival rates
are proportional to the covariate plus an 'intercept' term
for each site, plus a fixed amount of difference between the
two stages. Detection probabilities vary by stage.
-
Phi(site+BCOV+CCOV)p(s) - survival varies by site, binary
covariate (bcov) and the continous covariate (ccov) in 'parallel'.
With a continous
covariate, each nest with a different value for the covariate
can have a different survival rate, but the survival rates
are proportional to the covariate plus an 'intercept' term
for each site, plus a fixed amount of difference between the
two categories of the binary covariate. Detection probabilities vary by stage.
